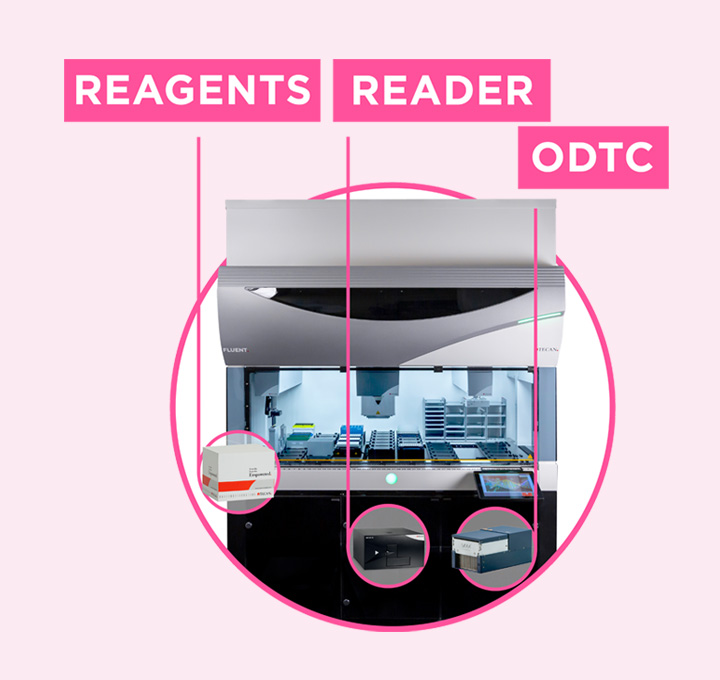
Freedom EVO NGS workstation
The Freedom EVO NGS workstation is equipped with innovative air displacement pipetting technology, using eight pipetting channels (1) to allow precise pipetting from 1,000 µl down to 0.5 µl. The workstation includes three Inheco CPAC devices (2) to keep reagents cool and provide optimal conditions for the enzymatic steps, an Inheco Thermoshake heated shaker (3), a 96-position Alpaqua 96S Super Magnet Plate magnetic plate separator (4) and a RoMa Arm (5) for efficient bead clean-up. In addition, the compact worktable provides storage space for up to 12 tip boxes (6), allowing longer unattended runs.
Precision: The Freedom EVO’s precision automation ensures the reproducibility necessary for high quality results, with a range of qualified sequencing-verified protocols available.
Flexibility: Don’t be constrained by batching requirements. Whether you need to process 6, 13, 19, 40 or even 96 samples, only run what you need, saving on reagents.
Simplicity: You don’t need to be an automation expert. With the simple touchscreen interface, you can get started right away, set up your runs quickly and be confident in your workflows.